Probability and Statistics
Est. read time: 3 minutes | Last updated: May 04, 2026 by John Gentile
Contents
import numpy as np
import matplotlib.pyplot as plt
from sympy import *
init_printing()
Distributions
There are several distributions relevant to detection and estimation. The Probability Density Function (PDF) gives the probability that a random vairable will take the value .
The main central moments are:
- Mean/Expectation:
- Variance:
Where is the expectation operator:
Other notations:
- The operator means “is distributed as”.
- means the distribution is used in the PDF.
Gaussian Random Variable
The Gaussian, or normal, distribution is defined with mean and variance of:
It is parameterized as:
With PDF implemented as:
def normal_pdf(x: np.ndarray, mu: float, sigma: float) -> np.ndarray:
return (1 / (sigma * np.sqrt(2 * np.pi))) * np.exp(-0.5 * ((x - mu) / sigma) ** 2)
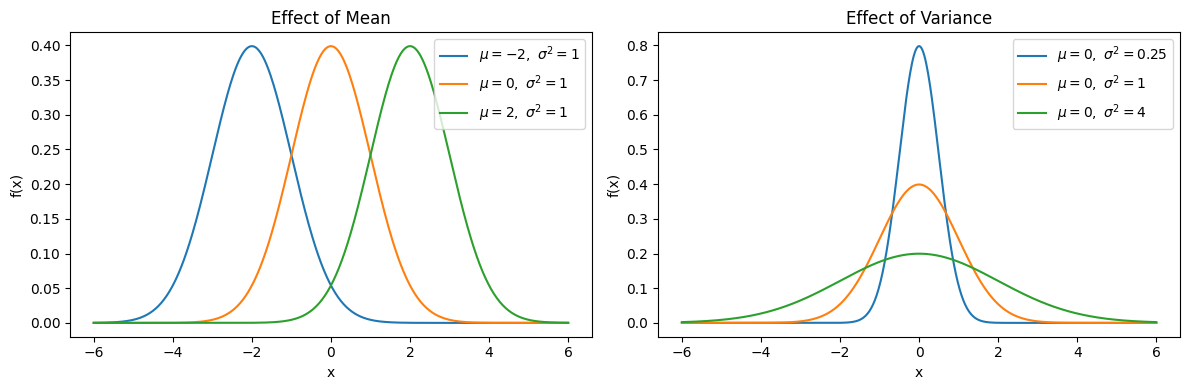
samples = np.linspace(-6, 6, 500)
fig, axes = plt.subplots(1, 2, figsize=(12, 4))
# Vary mean with fixed variance
for mu in [-2, 0, 2]:
sigma = 1
axes[0].plot(samples, normal_pdf(samples, mu, sigma), label=rf"$\mu={mu},\ \sigma^2={sigma**2}$")
axes[0].set_title("Effect of Mean")
axes[0].set_xlabel("x")
axes[0].set_ylabel("f(x)")
axes[0].legend()
# Vary variance with fixed mean
for sigma in [0.5, 1, 2]:
mu = 0
axes[1].plot(samples, normal_pdf(samples, mu, sigma), label=rf"$\mu={mu},\ \sigma^2={sigma**2}$")
axes[1].set_title("Effect of Variance")
axes[1].set_xlabel("x")
axes[1].set_ylabel("f(x)")
axes[1].legend()
plt.tight_layout()
plt.show()
Detection Theory
Usually called hypothesis testing, in DSP the simplest form of signal detection is shown as a binary hypothesis- the two hypotheses are commonly referred to as the null hypothesis (, signal is absent) and the alternative hypothesis (, signal is present)
Estimation Theory
Maximum Likelihood Estimation
Maximum Likelihood Estimation (MLE) is a method for estimating the parameters of a distribution given observed data. The idea is to choose the parameters that make the observed data most probable.
Given independent observations drawn from a distribution with parameter(s) , the likelihood function is the joint probability of the data:
In practice we work with the log-likelihood (since products become sums):
The MLE is the value that maximizes :
Example: MLE for the Gaussian Mean
If we assume with known , the log-likelihood as a function of is:
Setting and solving gives the familiar result:
The MLE of the mean is simply the sample mean.
np.random.seed(42)
# Generate samples from a known Gaussian
true_mu = 3.0
true_sigma = 1.5
N = 30
samples = np.random.normal(true_mu, true_sigma, N)
# Compute log-likelihood over a range of candidate mu values (sigma known)
mu_range = np.linspace(-1, 7, 500)
log_likelihood = np.array([
-N/2 * np.log(2 * np.pi * true_sigma**2)
- 1/(2 * true_sigma**2) * np.sum((samples - mu)**2)
for mu in mu_range
])
mu_ml = np.mean(samples)
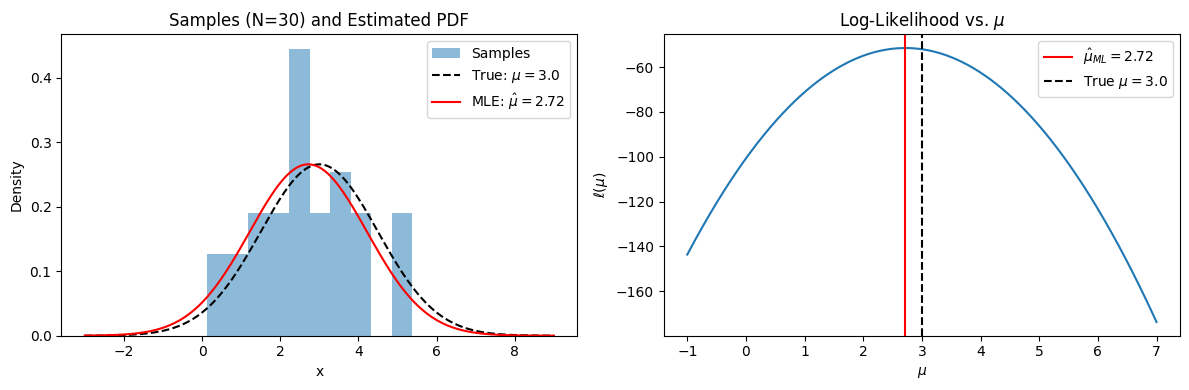
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(12, 4))
# Left: the observed samples and the true vs estimated distributions
ax1.hist(samples, bins=10, density=True, alpha=0.5, label="Samples")
x_plot = np.linspace(-3, 9, 300)
ax1.plot(x_plot,
(1 / (true_sigma * np.sqrt(2 * np.pi))) * np.exp(-0.5 * ((x_plot - true_mu) / true_sigma)**2),
"k--", label=rf"True: $\mu={true_mu}$")
ax1.plot(x_plot,
(1 / (true_sigma * np.sqrt(2 * np.pi))) * np.exp(-0.5 * ((x_plot - mu_ml) / true_sigma)**2),
"r-", label=rf"MLE: $\hat={mu_ml:.2f}$")
ax1.set_xlabel("x")
ax1.set_ylabel("Density")
ax1.set_title(f"Samples (N={N}) and Estimated PDF")
ax1.legend()
# Right: the log-likelihood curve
ax2.plot(mu_range, log_likelihood)
ax2.axvline(mu_ml, color="r", linestyle="-", label=rf"$\hat_={mu_ml:.2f}$")
ax2.axvline(true_mu, color="k", linestyle="--", label=rf"True $\mu={true_mu}$")
ax2.set_xlabel(r"$\mu$")
ax2.set_ylabel(r"$\ell(\mu)$")
ax2.set_title(r"Log-Likelihood vs. $\mu$")
ax2.legend()
plt.tight_layout()
plt.show()
Covariance Matrices
For a random vector with mean , the covariance matrix is:
where denotes the conjugate transpose. Each element of is the covariance between and :
The diagonal entries are the variances of each element.
Key Properties
- Hermitian: (symmetric for real data)
- Positive semi-definite: for all , meaning all eigenvalues
- Eigendecomposition: where is unitary and
Sample Covariance Matrix
In practice we estimate from observation snapshots :
(assuming zero-mean data, or after subtracting the sample mean). As , .
Relevance to Array Processing
In sensor array processing (e.g. radar, sonar, communications), is the snapshot vector across antenna elements. The covariance matrix encodes the spatial structure of all impinging signals and noise. Algorithms like MVDR beamforming and MUSIC operate directly on (or its estimate) to detect signals and estimate their directions of arrival.
np.random.seed(0)
# --- Simulate a uniform linear array (ULA) with M elements ---
M = 8 # number of array elements
N = 1000 # number of snapshots
d_lambda = 0.5 # element spacing in wavelengths
# Two signals arriving from different angles (degrees)
angles_deg = np.array([20, -35])
angles_rad = np.deg2rad(angles_deg)
signal_powers = np.array([10, 5]) # linear power
noise_power = 1.0
# Steering vectors: a(theta) = exp(j * 2pi * d * m * sin(theta))
m_idx = np.arange(M)
A = np.exp(1j * 2 * np.pi * d_lambda * np.outer(m_idx, np.sin(angles_rad))) # M x K
# Generate snapshots: x = A @ s + n
S = (np.sqrt(signal_powers / 2)[:, None]
* (np.random.randn(len(angles_deg), N) + 1j * np.random.randn(len(angles_deg), N)))
noise = np.sqrt(noise_power / 2) * (np.random.randn(M, N) + 1j * np.random.randn(M, N))
X = A @ S + noise
# --- Build sample covariance matrix ---
R_hat = (X @ X.conj().T) / N
# --- Verify key properties ---
print("Shape:", R_hat.shape)
print(f"Hermitian: {np.allclose(R_hat, R_hat.conj().T)}")
eigenvalues = np.linalg.eigvalsh(R_hat)
print(f"Eigenvalues (sorted): {np.sort(eigenvalues)[::-1].round(3)}")
print(f"All eigenvalues >= 0: {np.all(eigenvalues >= -1e-10)}")
print(f"Number of dominant eigenvalues: {np.sum(eigenvalues > 2 * noise_power)} (expect {len(angles_deg)} signals)")
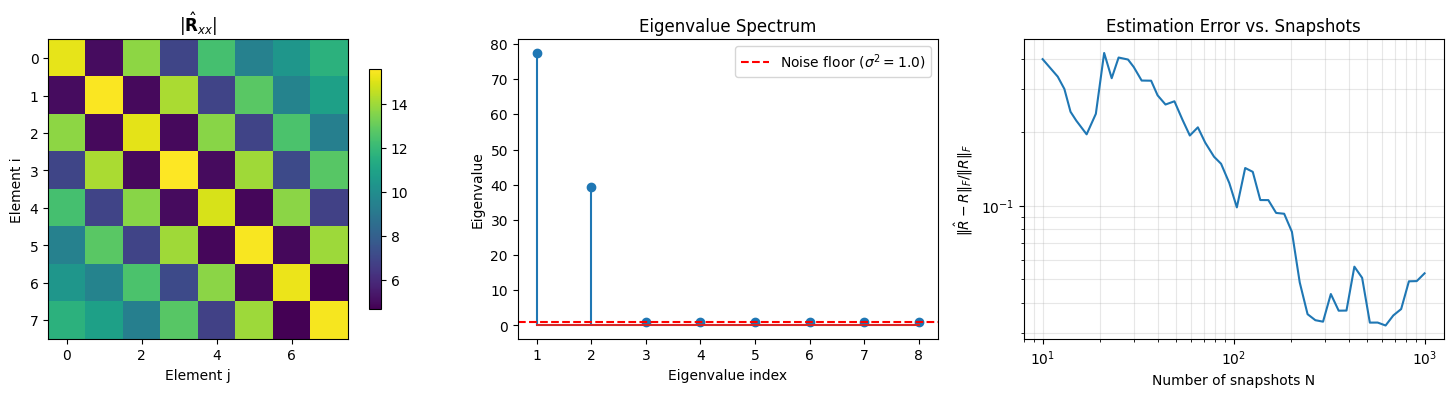
# --- Visualize ---
fig, axes = plt.subplots(1, 3, figsize=(15, 4))
# 1) Covariance matrix magnitude
im = axes[0].imshow(np.abs(R_hat), cmap="viridis")
axes[0].set_title(r"$|\hat{\mathbf{R}}_{xx}|$")
axes[0].set_xlabel("Element j")
axes[0].set_ylabel("Element i")
fig.colorbar(im, ax=axes[0], shrink=0.8)
# 2) Eigenvalue spectrum
axes[1].stem(range(1, M + 1), np.sort(eigenvalues)[::-1])
axes[1].axhline(noise_power, color="r", linestyle="--", label=rf"Noise floor ($\sigma^2={noise_power}$)")
axes[1].set_xlabel("Eigenvalue index")
axes[1].set_ylabel("Eigenvalue")
axes[1].set_title("Eigenvalue Spectrum")
axes[1].legend()
# 3) Convergence: sample covariance error vs number of snapshots
# True covariance: R = A @ diag(powers) @ A^H + sigma^2 I
R_true = A @ np.diag(signal_powers) @ A.conj().T + noise_power * np.eye(M)
n_values = np.logspace(1, np.log10(N), 50).astype(int)
errors = []
for n in n_values:
R_n = (X[:, :n] @ X[:, :n].conj().T) / n
errors.append(np.linalg.norm(R_n - R_true, "fro") / np.linalg.norm(R_true, "fro"))
axes[2].loglog(n_values, errors)
axes[2].set_xlabel("Number of snapshots N")
axes[2].set_ylabel(r"$\|\hat{R} - R\|_F / \|R\|_F$")
axes[2].set_title("Estimation Error vs. Snapshots")
axes[2].grid(True, which="both", alpha=0.3)
plt.tight_layout()
plt.show()
Shape: (8, 8) Hermitian: True Eigenvalues (sorted): [77.446 39.351 1.126 1.046 0.989 0.961 0.92 0.862] All eigenvalues >= 0: True Number of dominant eigenvalues: 2 (expect 2 signals)